1.课题组组长简介
常啸,医学人工智能与大数据学院教授、转化基因组学课题组长,国家级海外青年人才项目评审,山东省泰山学者青年专家。博士毕业于中国科学院上海生命科学学院。曾任职于美国南加州大学、费城儿童医院。目前主要从事人类遗传疾病和生物信息方向的研究,代表论文以第一作者或通讯作者发表在Cell Research, The Innovation, Nature Communications (四篇), Journal of the National Cancer Institute (两篇), Journal of Allergy and Clinical Immunology (四篇),Moleculary Psychiatry, Biological Psychiatry 和 JAACAP等国际高影响力杂志。
2. 课题组研究方向
u方向一:生物医学AI算法与工具开发
研发深度学习与自然语言处理驱动的算法,推动遗传变异检测与功能注释工具的革新,提升疾病相关基因解析的精准性。
u方向二:AI驱动的东亚人群疾病易感性研究
利用深度学习与迁移学习技术,解析东亚人群特异性遗传变异的疾病关联性,开发AI驱动的疾病风险预测模型。
u方向三:AI赋能的复杂疾病多组学机制解析
基于深度学习、图神经网络等技术,整合基因组、转录组、代谢组等多模态数据,解析疾病遗传调控网络。
3. 学术论文
近三年来,课题组共发表通讯/一作论文 20 篇, 其中5年影响因子大于10分文章 8 篇,一区论文11篇。
代表性论文:
(1) Ziang Meng#, Yingchao Song#, Yue Jiang, Xianjin Wang, Yang Zou, Xingyuan Li, Hakon Hakonarson*, Xiao Chang*; Transformer-based deep learning enhances discovery in migraine GWAS; Nature Communications, 2025 Dec 10, 16(1): 11023. (期刊论文;IF=15.7)
(2) Lili Zhi#, Qiwen Zheng#, Yue Jiang#, Lu Yu, Linzehao Li, Yingchao Song, Bichen Peng, Chumeng Zhang, Hengxuan Jiang, Ren Li, Frank Mentch, Joseph Glessner, Peilin Jia, Hua Tang*, Hakon Hakonarson*, Xiao Chang*; Multi-trait genetic analysis of asthma and eosinophils uncovers pleiotropic loci in East Asians; Nature Communications, 2025, 16: 5081. (期刊论文;IF=15.7)
(3) Xiao Chang*, Huiqi Qu, Yichuan Liu, Joseph Glessner, Hakon Hakonarson*; A Protective Role of Low Polygenic Risk Score in Healthy Individuals Carrying Attention-Deficit/Hyperactivity Disorder-Associated Copy Number Variations; Biological Psychiatry, 2024 May 1, 95(9): 881-887. (期刊论文;IF=9.6)
(4) Xiao Chang*, Hui-Qi Qu, Yichuan Liu, Joseph T Glessner, Hakon Hakonarson; Mitochondrial DNA Haplogroup K Is Protective Against Autism Spectrum Disorder Risk in Populations of European Ancestry; Journal of the American Academy of Child and Adolescent Psychiatry, 2024 Aug, 63(8): 835-844. (期刊论文;IF=9.2)
(5) Yunhai Wei#, Ziang Meng#, Xianjin Wang, Yue Jiang, Huanxin Ding, Bichen Peng, Yingchao Song, Min Gao, Guangyong Zhang, Nan Zhang, Xiao Chang*; Transformer-based InsightGWAS improves GERD genetic discovery via pretraining on GWAS for major depressive disorder; Communications Biology, 2026 Jan 05, 9: 2. (期刊论文;IF=5.9)
(6) Yingchao Song#, Linzehao Li#, Yue Jiang, Bichen Peng, Hengxuan Jiang, Zhen Chao, Xiao Chang*; Multitrait Genetic Analysis Identifies Novel Pleiotropic Loci for Depression and Schizophrenia in East Asians; Schizophrenia Bulletin, 2025 May 8, 51(3): 684-695. (期刊论文;IF=5.3)
(7) Huanxin Ding#, Yue Jiang#, Qing Sun, Yingchao Song, Shuohui Dong, Qian Xu, Linzehao Li, Chuxuan Liu, Bingjun Li, Hengxuan Jiang, Bichen Peng, Shi Peng, Chumeng Zhang, Jiankang Zhu, Mingwei Zhong, Guangyong Zhang*, Xiao Chang*; Integrating genetics and transcriptomics to characterize shared mechanisms in digestive diseases and psychiatric disorders; Communications Biology, 2025, 8: 47. (期刊论文;IF=5.2)
(8) Xiao Chang*, Michael March, Frank Mentch, Huiqi Qu, Yichuan Liu, Joseph Glessner, Patrick Sleiman, Hakon Hakonarson*; Genetic architecture of asthma in African American patients; Journal of Allergy and Clinical Immunology, 2023 Apr, 151(4): 1132-1136. (期刊论文;IF=11.4)
(9) Christopher J Cardinale#, Xiao Chang#, Zhi Wei, Hui-Qi Qu, Jonathan P Bradfield, Constantin Polychronakos, Hakon Hakonarson*; Genome-wide association study of the age of onset of type 1 diabetes reveals HTATIP2 as a novel T cell regulator; Frontiers in Immunology, 2023 Feb 1, 14: 1101488. (期刊论文;IF=5.7)
(10) Xiao Chang*, Michael March, Frank Mentch, Huiqi Qu, Yichuan Liu, Joseph Glessner, Patrick Sleiman, Hakon Hakonarson*; Genetic architecture of asthma in African American Patients; Journal of Allergy and Clinical Immunology, 2022 Sep 8. (期刊论文;IF=14.2)
(11) Xiao Chang, Yichuan Liu, Joseph Glessner, Cuiping Hou, Huiqi Qu, Kenny Nguyen, Patrick Sleiman, Lobin Lee, Sharon J Diskin, John M Maris, Hakon Hakonarson*; Identification of Mitochondrial DNA Variants Associated With Risk of Neuroblastoma, JNCI: Journal of the National Cancer Institute, 2022, 114(6): 910-913. (期刊论文;IF=10.3)
(12) Xiao Chang, Michael March, Frank Mentch, Kenny Nguyen, Joseph Glessner, Huiqi Qu, Yichuan Liu, Glen Furuta, Seema Aceves, Nirmala Gonsalves, Kari Nadeau, Antonella Cianferoni, Jonathan Spergel, Patrick Sleiman, Hakon Hakonarson*; A genome-wide association meta-analysis identifies new eosinophilic esophagitis loci, Journal of Allergy and Clinical Immunology, 2022, 149(3): 988-998. (期刊论文; IF=14.2)
(13) Xiao Chang; Marina Bakay; Yichuan Liu; Joseph Glessner; Komal S Rathi; Cuiping Hou; Huiqi Qu; Zalman Vaksman; Kenny Nguyen; Patrick M A Sleiman; Sharon J Diskin; John M Maris; Hakon Hakonarson*; Mitochondrial DNA haplogroups and susceptibility to neuroblastoma, JNCI: Journal of the National Cancer Institute, 2020, 112(12): 1259-1266. (期刊论文;IF=10.3)
(14) Stanislaw J. Gabryszewski#; Xiao Chang#; Jesse W. Dudley; Frank Mentch; Michael March; John H. Holmes; Jason Moore; Robert W. Grundmeier; Hakon Hakonarson; David A. Hill*; Unsupervised modeling and genome-wide association identify novel features of allergic march trajectories, Journal of Allergy and Clinical Immunology, 2021, 147(2): 677-685. (期刊论文;IF=14.2)
(15) Xiao Chang#, Yun Li#, Kenny Nguyen, Huiqi Qu, Yichuan Liu, Joseph Glessner, Patrick MA Sleiman, Hakon Hakonarson*; Genetic correlations between COVID-19 and a variety of traits and diseases, The Innovation, 2021, 2(2): 100112. (期刊论文;IF=32.1)
(16) Xiao Chang; Yan Zhao; Cuiping Hou; Joseph Glessner; Lee McDaniel; Maura A. Diamond; Kelly Thomas; Jin Li; Zhi Wei; Yichuan Liu; Yiran Guo; Frank D. Mentch; Haijun Qiu; Cecilia Kim; Perry Evans; Zalman Vaksman; Sharon J. Diskin; Edward F. Attiyeh; Patrick Sleiman; John M. Maris*; Hakon Hakonarson*; Common variants in MMP20 at 11q22.2 predispose to 11q deletion and neuroblastoma risk, Nature Communications, 2017, 8(1): 1-7. (期刊论文;IF=16.6)
(17) Yi Zhong#; Xiao Chang#; Xing-Jun Cao#; Yan Zhang; Huajun Zheng; Yongzhang Zhu; Chengsong Cai; Zelin Cui; Yunyi Zhang; Yuan-Yuan Li; Xiu-Gao Jiang; Guo-Ping Zhao; Shengyue Wang*; Yixue Li*; Rong Zeng*; Xuan Li*; Xiao-Kui Guo*; Comparative proteogenomic analysis of the Leptospira interrogans virulence-attenuated strain IPAV against the pathogenic strain 56601, Cell Research, 2011, 21(8): 1210-1229. (期刊论文;IF=44.1)
4. 研究成果展示
uGWAS人工智能方法学开发
基于Transformer架构的偏头痛易感基因智能筛选
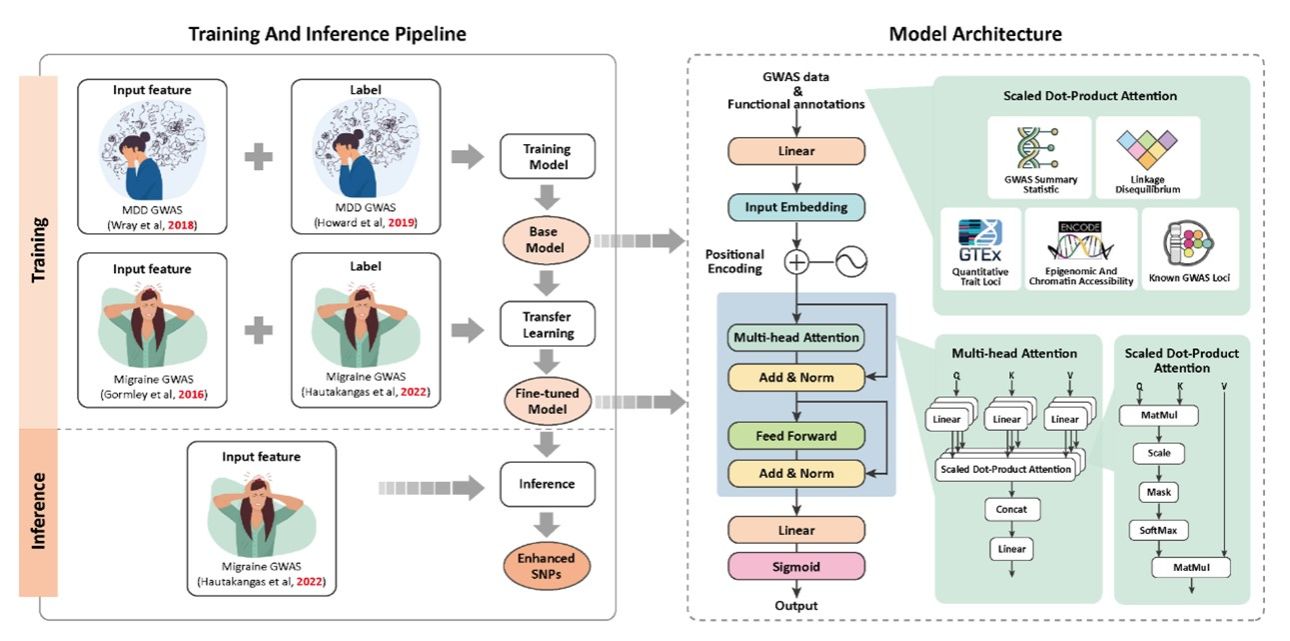
u跨表型GWAS数据分析
多表型联合GWAS分析探索东亚人群中哮喘的潜在靶点基因
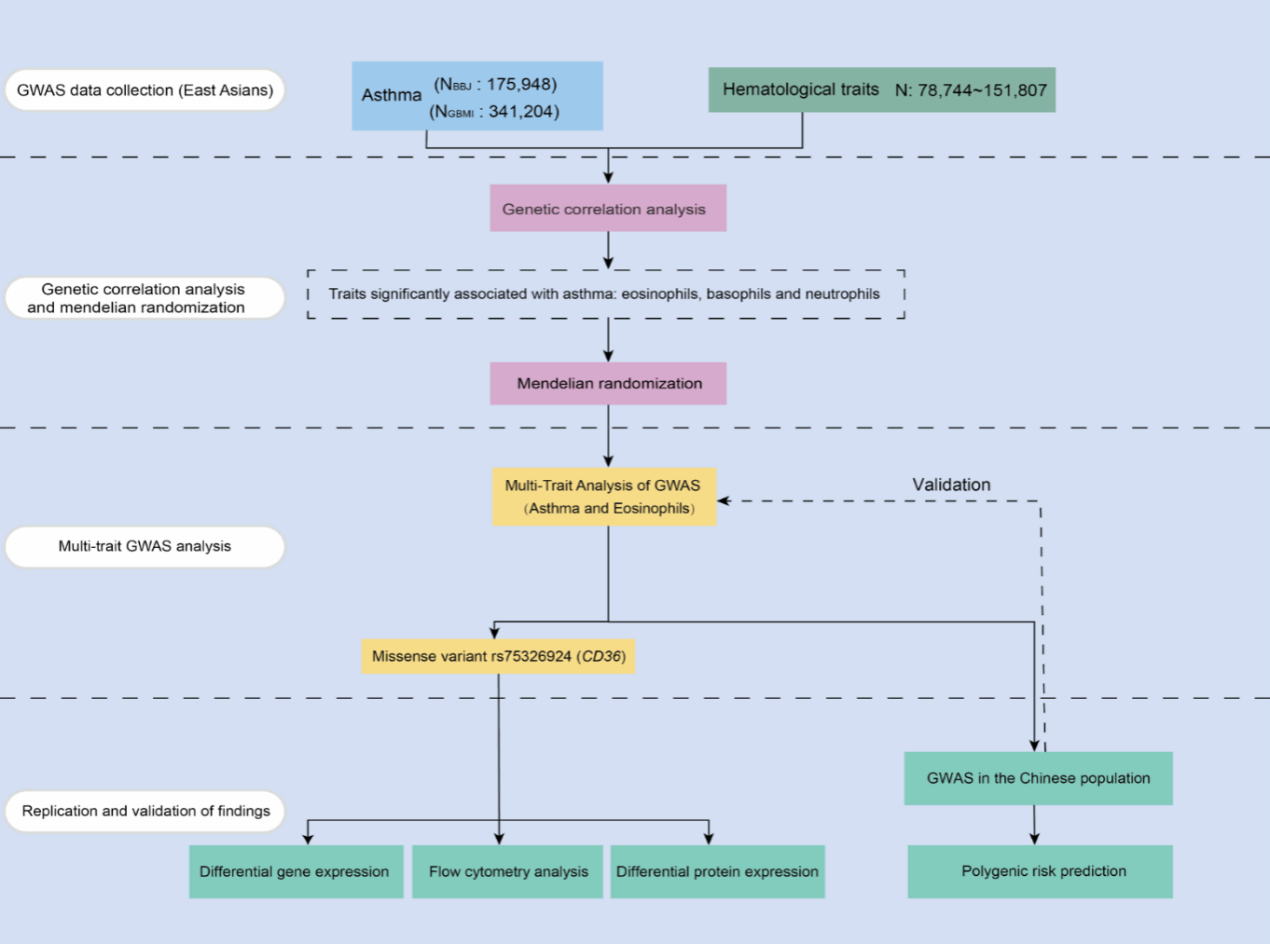
多表型联合GWAS分析探究精神疾病与消化道疾病遗传共病机制
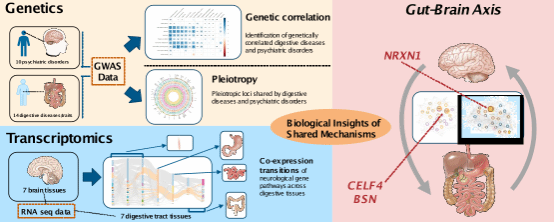
多表型联合GWAS分析挖掘东亚人群中抑郁症和精神分裂症的多效基因位点
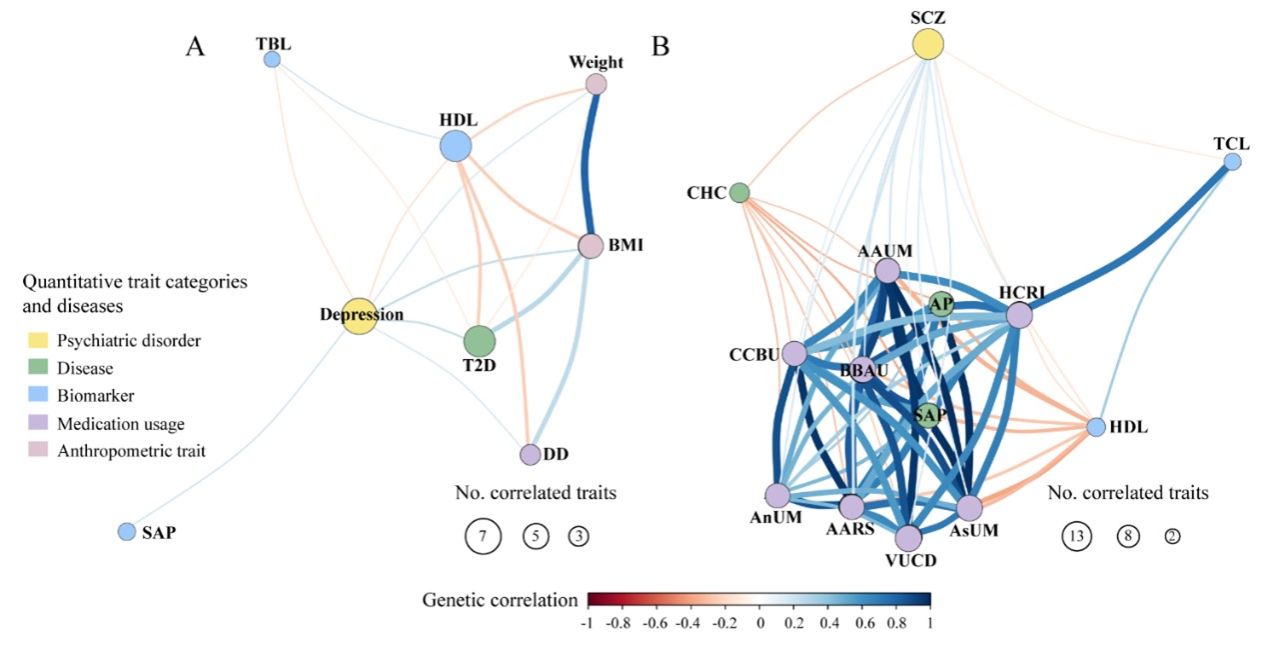
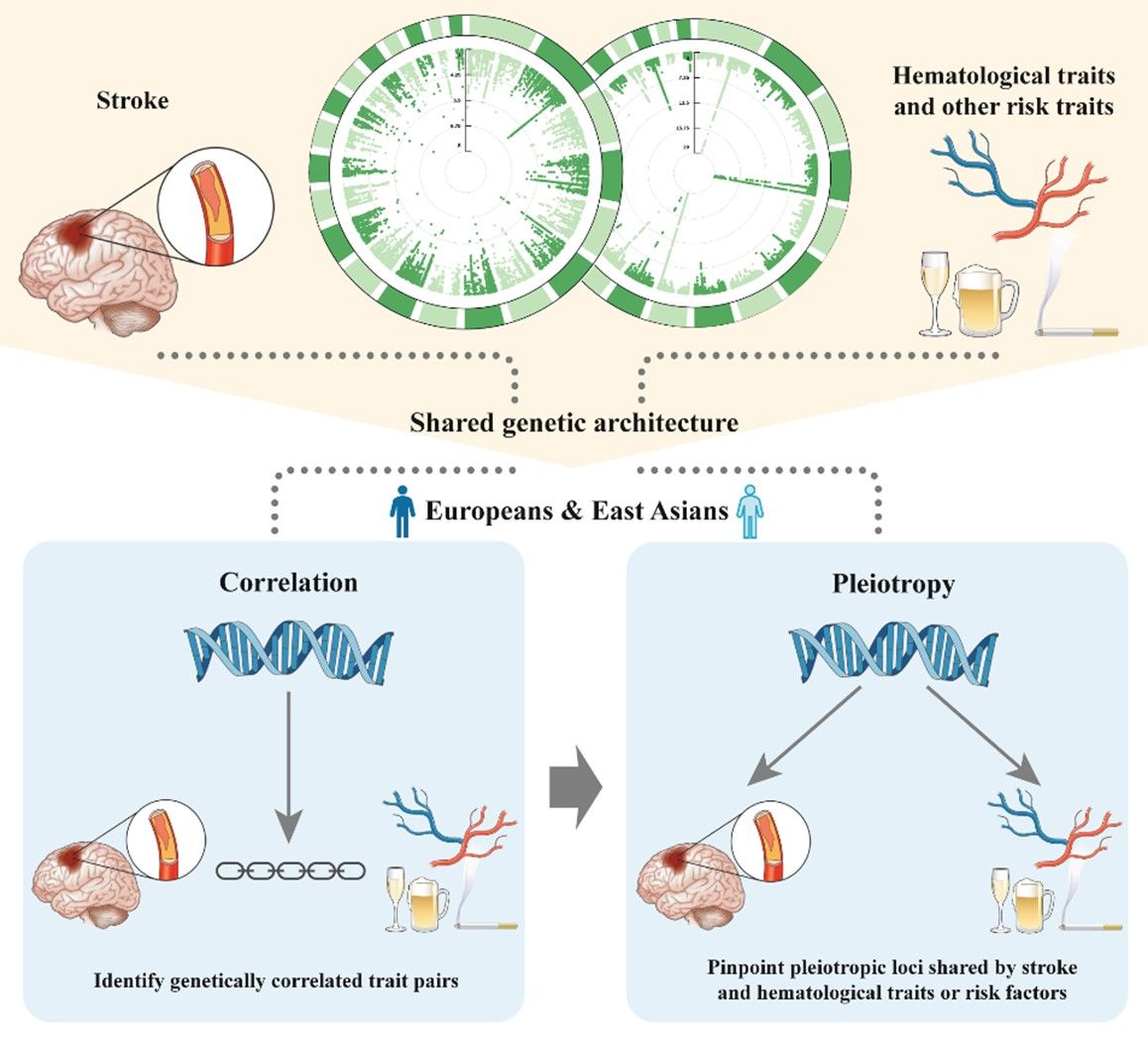
多表型联合GWAS分析探索中风与血液风险因素的多效基因
5. 科研项目
u国家自然科学基金面上项目(项目号32270661,54万元,常啸);
u山东省优秀青年科学基金项目(海外,60万元,常啸);
u中国博士后科学基金第76批面上资助(项目号2024M761870,8万元,宋颖超);
u山东省博士后创新项目(项目号SDCX-ZG-202400042,3万元,宋颖超 )。
6. 人才培养
团队的研究生、本科生获奖数项
u第十届全国大学生统计建模大赛
获奖名次:山东赛区研究生组三等奖
作品名称:基于深度神经网络的重症CCOVID-19全基因组位点预测与分析
参赛队员:彭碧晨、姜越、叶伟逸
指导教师:常啸
u第十一届全国大学生医学创新大赛暨"一带一路"国际竞赛
获奖名次:三等奖
作品名称:解码偏头痛遗传密钥:基于Transformer模型的深度学习驱动全基因组关联研究新突破
参赛队员:李玟、张楚萌、王丽轩、王越、雷雨昕
指导教师:常啸
u2025年"挑战杯"大学生课外学术科技作品竞赛
获奖名次:二等奖
作品名称:基于Transformer模型的深度学习助力偏头痛全基因相关联研究的新发现
参赛队员:李玟、张楚萌、王越、孙紫然、雷雨昕、赵俊凯
指导教师:常啸
u2025年"挑战杯"大学生课外学术科技作品竞赛
获奖名次:二等奖
作品名称:通过多性状GWAS分析探索与COVID-19严重程度和免疫反应相关的遗传位点
参赛队员:王越、程静、李玟、张楚萌、孙紫然
指导教师:常啸

7. 团队建设
u团队合照

u团队成员

8. 长期招聘
1.招聘博士后:要求生物医学工程、电子信息、计算机科学与技术等相关专业背景。具有深度学习、基因组学数据分析或工具开发经验者优先。
2.招聘研究生或访问学生:对生物信息学与AI交叉研究有浓厚兴趣;能保证连续学习时间不少于1年;有相关研究经历者优先。
有意向者可发简历到邮箱:ycsong@sdfmu.edu.cn,将会在3个工作日内回复。

